|
|
Witmer, L.M., Ridgely, R.C., Mayle, H., and D. Adams. 2004. The best of both worlds: integrating CT and MR. RT image 17(32):16-19.
Pigs (Sus scrofa) are known far and wide as the wise and noble animals of the barnyard. Of all modern species in the family Suidae, S. scrofa has the widest geographical range, which is due in no small part to their domestication by humans ~4900 years ago. Subsequently, pigs have been introduced virtually everywhere that humans have gone, and have been selectively bred to produce a wide variety of breeds the number of which rivals that of the domestic dog.
In the biomedical field, pigs are the animal of choice for modeling human anatomical systems because of their availability and proximity in size to humans. These wide ranging applications include drug testing and organ transplants, as well as understanding disorders of the temporomandibular joint, degenerative joint disease, and the production of joint prostheses.

About the Species
This specimen is a left hind limb, removed from a female Hanford mini-pig that had previously been euthanized as part of an experimental protocol by other Ohio University researchers. The goal of the current study was to produce a 3D model of both the boney and soft tissues of the knee to facilitate biomechanical study of the joint. To image the relevant tissues, it was necessary to collect data using both X-ray CT and magnetic resonance imaging (MRI).
To allow the registration and integration of these different types of data, it was necessary to fix the limb in a rigid medium to inhibit movement about the knee so that it would be in the same relative position in the different scans. This was accomplished using an expandable, insulating foam, which is of extremely low density and quite rigid when cured (Figures 1 and 2).
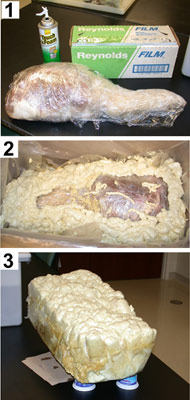 |
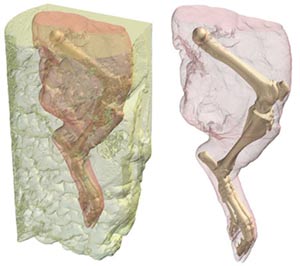 |
Figure 1 |
Figure 2 |

About this Specimen
The limb, now embedded in a foam block, was then scanned in CT and MRI at O'Bleness Memorial Hospital in Athens, Ohio. The CT scans were performed on a GE HiSpeed Fx-i Helical CT scanner (120.0 kV, 100.0 mA, axial slice thickness of 1 mm). MR imaging was performed on a 1.0T GE Signa Short-Bore MRI system (T1, axial and sagittal slice thickness of 3 mm). The entire hindlimb was CT scanned, but the MR field of view was limited to the knee region. The scan data were then imported into a computer workstation, and analyzed using Amira 3.1 (TGS, Inc., San Diego). Because the 3mm slice thicknesses of the MRI datasets are rather coarse for generating 3D models, the axial and sagittal MR datasets were combined and resampled to produce a composite volume of 1mm intervals (Figure 3). |
|
3D modeling and registration
Using this composite MR dataset, the medial and lateral menisci of the knee were "segmented", or selected, from the MR data, as were the knee ligaments (not shown). The bones of the hindlimb were segmented from the CT data, and rendered in 3D (Figure 4). Because the bone and menisci are from two separate data sets (CT and MR, respectively), it was necessary to register the two datasets in order to put them into the proper anatomical position. Fortunately, Amira 3.1 has an auto-registration function allowing the user to automatically register the MR and CT datasets once they have been grossly aligned. After the two datasets have been registered, the resulting registration parameters can then be applied to the 3D model of the menisci that had already generated, and it slides right into the knee (Figure 4). |
|
Fusion
While 3D modeling of CT and MR data is extremely useful, at times it is still necessary to use the raw slice data, for example when examining the relationships between bone and soft tissues in the skull. Even with the MR and CT data registered, having two separate slice volumes is cumbersome and thwarts integrating the information from each set.
To produce a single dataset from separate CT and MR volumes, we merged the two datasets. Since both datasets provide at least some information on similar structures (muscle, for example), we stripped away the redundant data. In other words, we removed all of the grey values that were less dense than bone from the CT volume (e.g., soft tissue, air), and from the MR volume we removed anything below the soft-tissue threshold (Figure 5). |
|
We then resampled, or resliced, the MR data set so that each slice in the MR volume corresponded directly to a slice at the same position in the CT dataset. Once this was done, we were able to merge the two datasets, which essentially blends a given CT slice with the corresponding MR slice. The result is a single dataset with bone from CT and the soft tissue from the MR.

About the
Scan
Literature
Witmer, L.M., Ridgely, R.C., Mayle, H., and D. Adams. 2004. The best of both worlds: integrating CT and MR. RT image 17(32):16-19
Links
Sus scrofa on the Animal Diversity Web (Univ. of Michigan Museum of Zoology)

Literature
& Links
MR/CT fusion movies:
Specimen/foam block movie: |
Knee with translating MR slice: |
 |
 |

Additional Imagery
 |
|

